HKUST Researchers Make Groundbreaking Discovery in RNA Silencing Mechanisms for Gene Regulation and Disease Insight
A research team from The Hong Kong University of Science and Technology (HKUST) has made significant strides in understanding the molecular mechanisms involved in RNA silencing. The study details how the human enzyme DICER conducts precise processing of microRNAs (miRNAs), which could advance research in gene regulation and provide insights into potential mechanisms related to cancer, immune disorders, and genetic diseases.
Research Team and Publication
The investigation was led by Professor Tuan Anh Nguyen, an Associate Professor in the Division of Life Science at HKUST, alongside PhD students Minh Khoa Ngo and Cong Truc Le. Their findings have been published in the academic journal Nature. The paper, titled “DICER cleavage fidelity is governed by 5′-end binding pockets,” extensively discusses the results of their research.
Understanding RNA and DICER Function
The human genome operates through messenger RNAs, which relay genetic instructions. RNA molecules, usually consisting of ribonucleotides (A, U, G, and C), are crucial for several cellular processes, including protein synthesis and gene regulation. DICER functions as “molecular scissors,” cleaving double-stranded RNA precursors into short regulatory RNAs that are incorporated into the RNA-induced silencing complex (RISC). This complex aids cells in recognizing and eliminating faulty genetic messages.
Research Methodology
Researchers have long aimed to clarify DICER’s accuracy in RNA cleavage. By employing advanced biochemical methodologies and structural biology techniques, including high-resolution cryoelectron microscopy (cryo-EM), the HKUST team was able to observe DICER-RNA interactions at the atomic level. According to Minh Khoa Ngo, cryo-EM offered insights into the intricate interactions between RNA substrates and DICER, revealing the dynamic adjustments the enzyme undergoes during the processing of various RNA sequences.
Key Findings
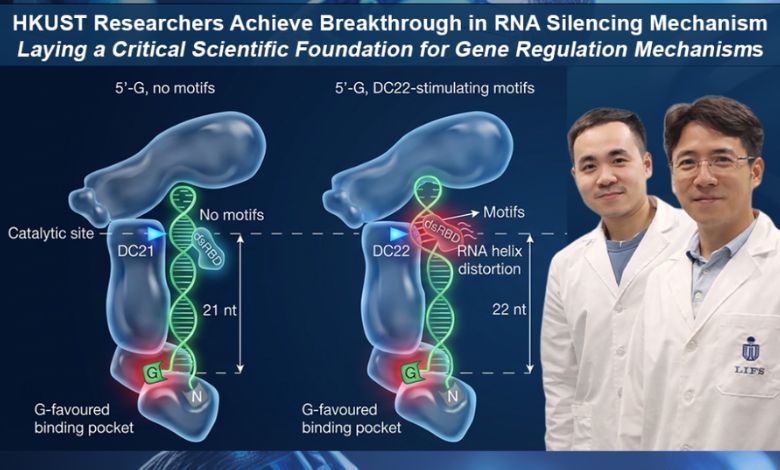
The findings suggest that before cleavage, DICER modifies its conformation to ensure correct alignment of RNA substrates. The enzyme employs distinct structural features within specific binding pockets to accurately position the RNA prior to entering a “cleavage-ready” state.
Implications of the Discovery
Professor Nguyen noted that DICER can pinpoint specific cleavage sites on RNA with single nucleotide resolution, thereby preserving the accuracy of genetic messaging. The study discovered both a U-favored and a newly identified G-favored 5′-end binding pocket, indicating a dual-pocket mechanism that positions cleavage. This revelation contributes to a deeper understanding of how DICER processes diverse RNA substrates.
Future Potential
This discovery has implications that extend beyond fundamental biology, as it reveals how DICER assimilates RNA characteristics to maintain cleavage fidelity. Such insights could facilitate the advancement of RNA-based therapeutics, enhance gene-silencing techniques, and shed light on genetic disorders associated with DICER.
(Source: HKUST)




